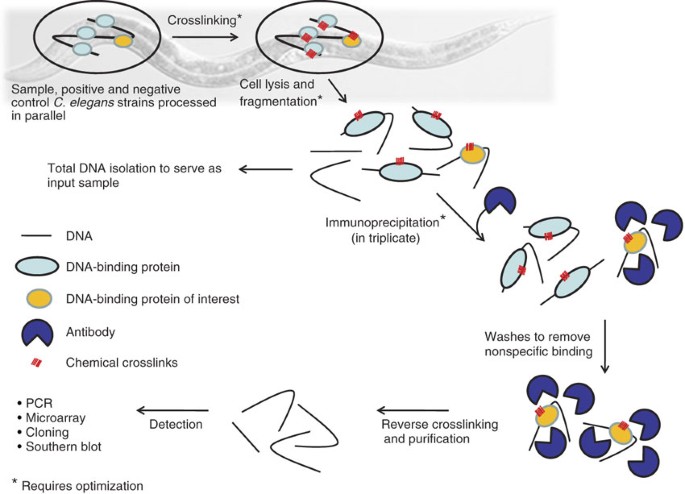
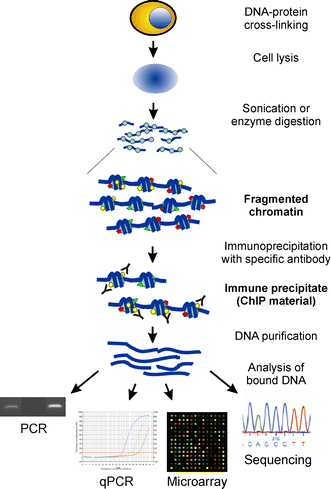
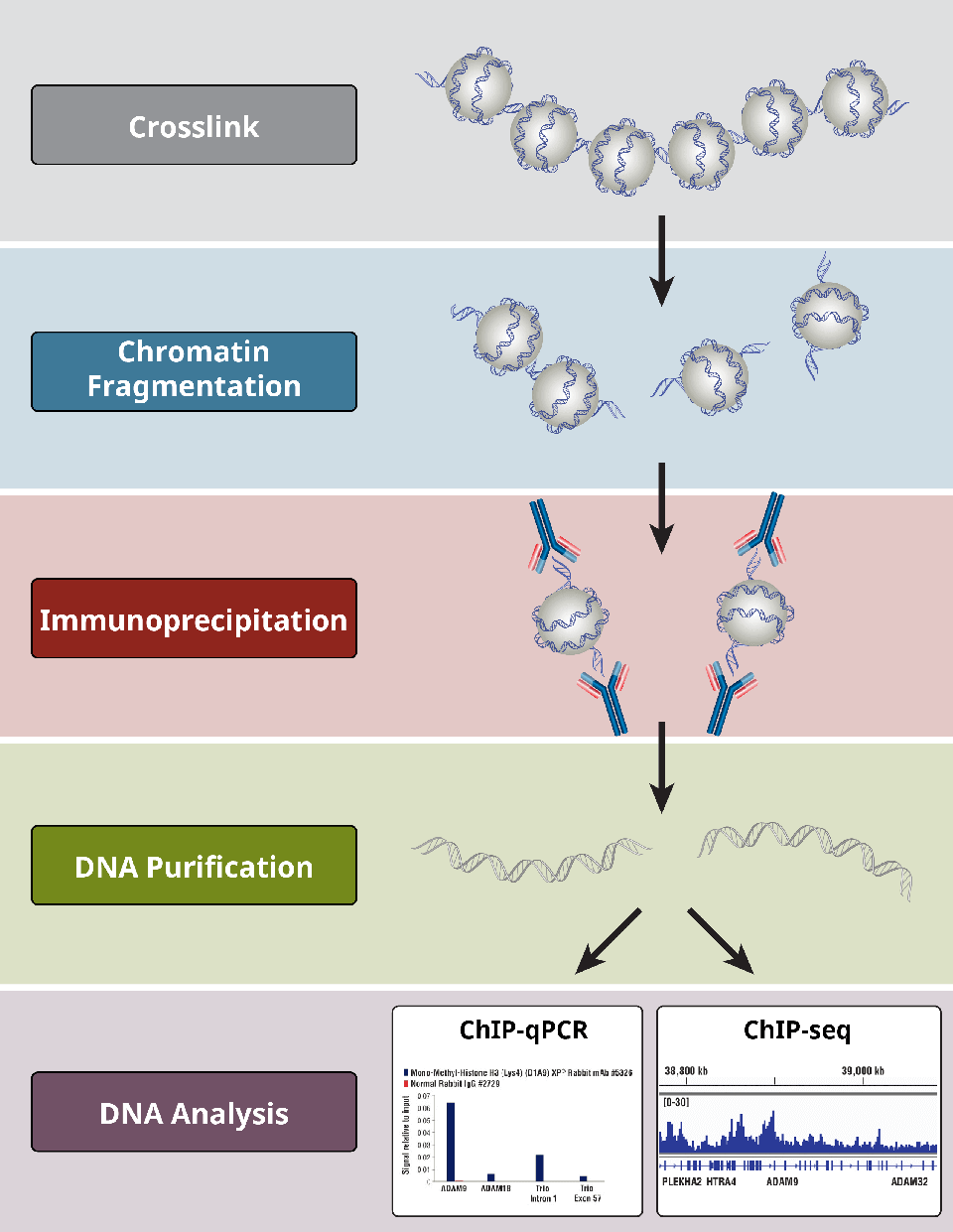
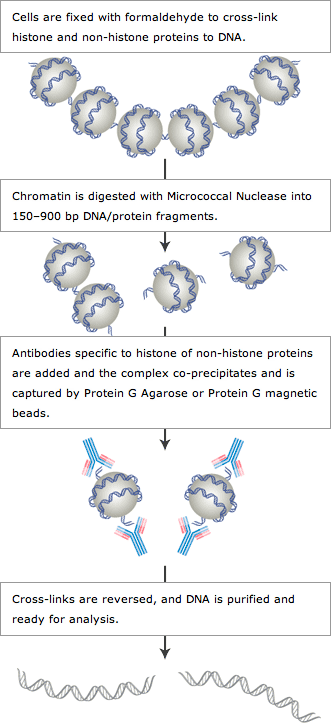
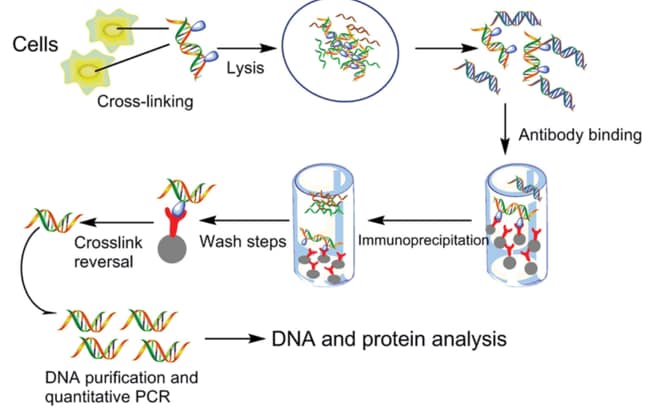
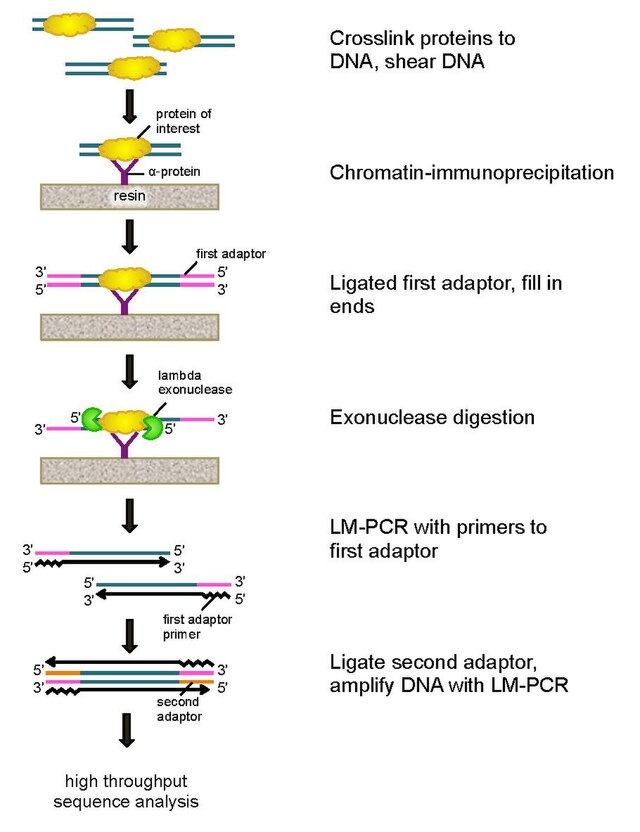
ChIP-on-chip protocol for genome-wide analysis of transcription factor binding in Drosophila melanogaster embryos | Nature Protocols

Protocol for fractionation-assisted native ChIP (fanChIP) to capture protein-protein/DNA interactions on chromatin - ScienceDirect
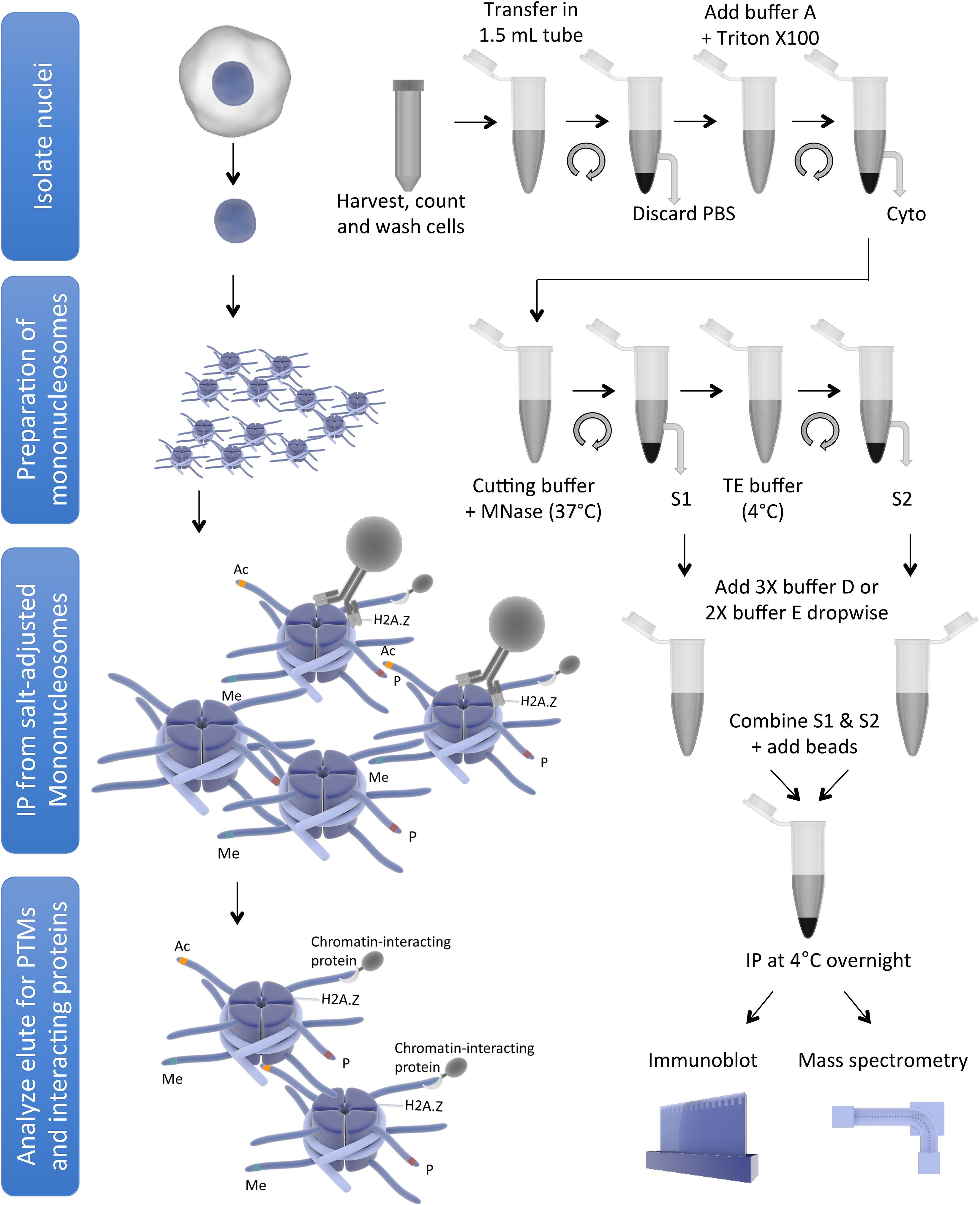
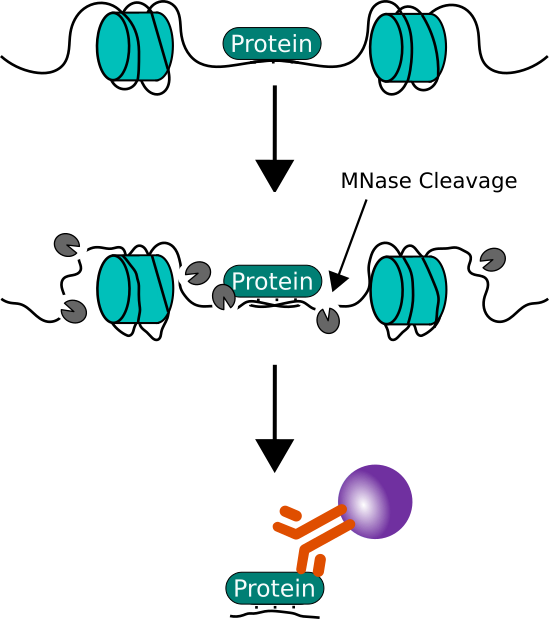
Frontiers | The Use of Mononucleosome Immunoprecipitation for Analysis of Combinatorial Histone Post-translational Modifications and Purification of Nucleosome-Interacting Proteins

Long-term association of a transcription factor with its chromatin binding site can stabilize gene expression and cell fate commitment | PNAS

Reverse Chromatin Immunoprecipitation (R-ChIP) enables investigation of the upstream regulators of plant genes | Communications Biology
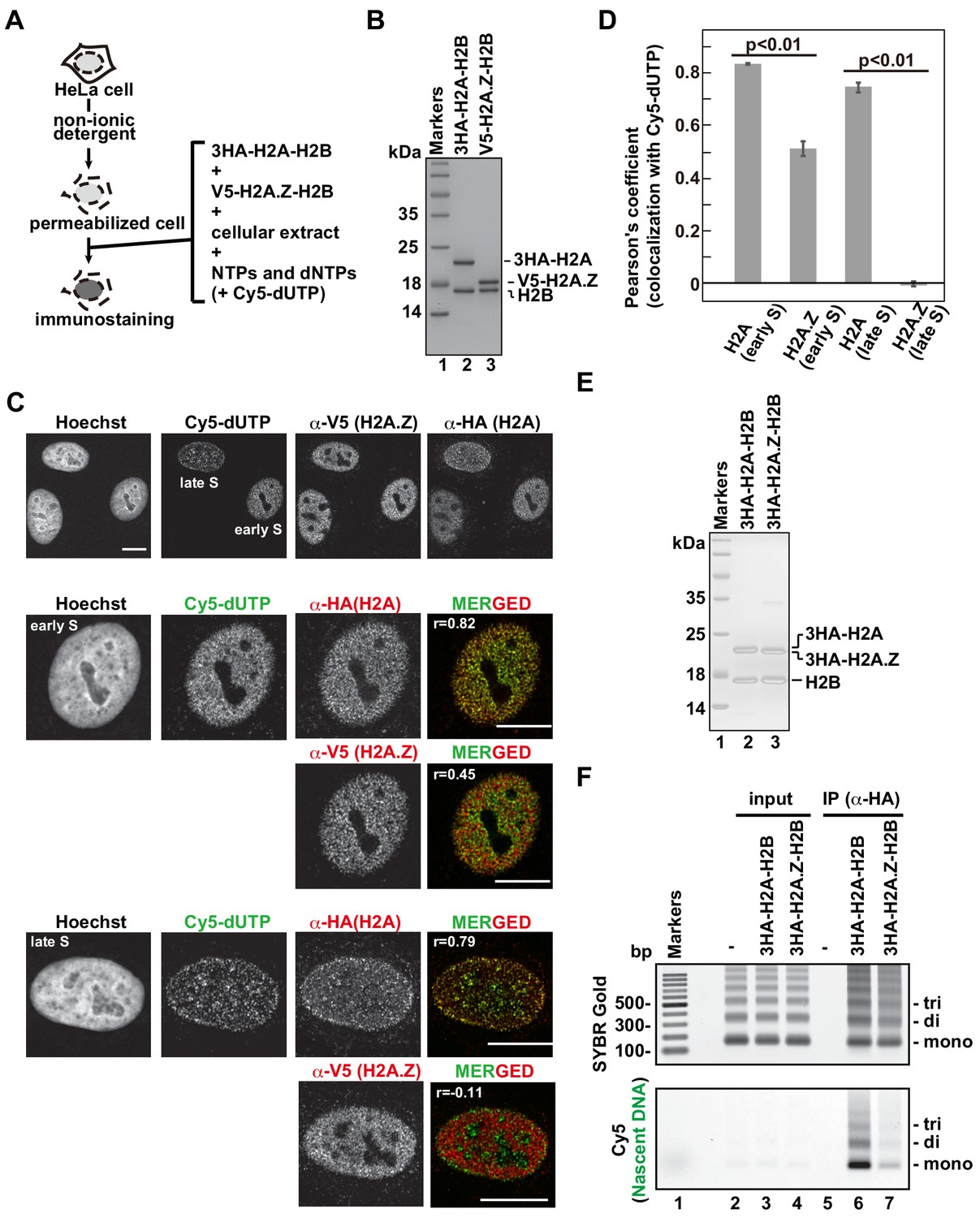
Chromatin structure-dependent histone incorporation revealed by a genome-wide deposition assay | eLife

Chromatin binding assays. (A) H1 chimeras are stringently chromatin... | Download Scientific Diagram
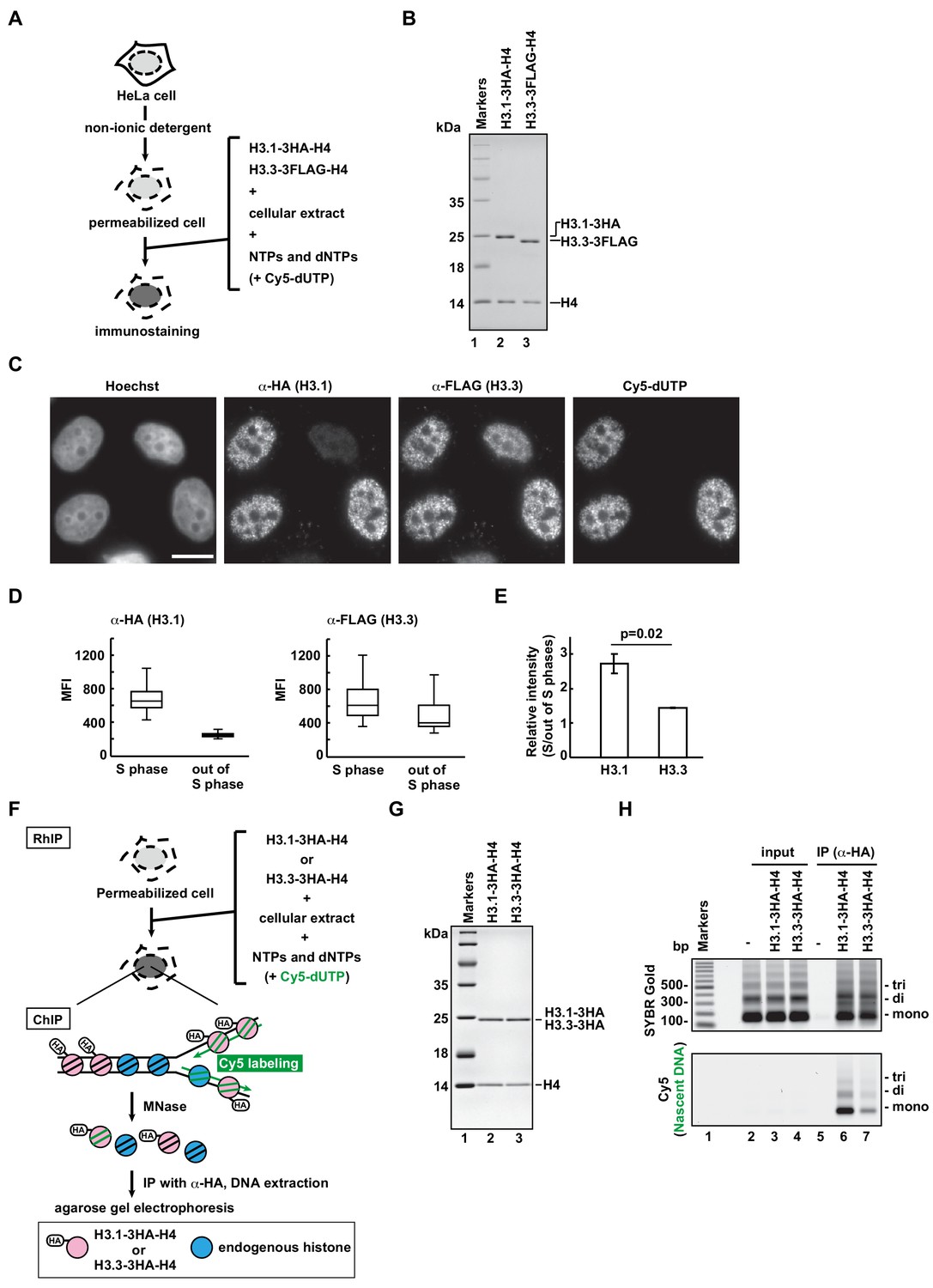
Chromatin structure-dependent histone incorporation revealed by a genome-wide deposition assay | eLife

An optimized two-step chromatin immunoprecipitation protocol to quantify the associations of two separate proteins and their common target DNA - ScienceDirect

Chromatin Immunoprecipitation Assay to Identify Genomic Binding Sites of Regulatory Factors | SpringerLink